Protein microarrays have emerged as an effective tool for efficiently analyzing various protein interactions. This advanced method provides valuable insights into cellular functions, disease mechanisms, and drug interactions, ultimately accelerating scientific discoveries in various fields, including oncology and infectious diseases.
This blog post will explore protein microarrays, their technology, applications, and potential impacts on biomedical research.
Understanding Protein Microarrays
Protein microarrays are small-scale versions of traditional laboratory assays, like ELISA. They consist of thousands of unique proteins fixed onto a solid surface, such as a glass slide or a microplate. Each protein spot works as a capture molecule. Protein microarrays have been used for various purposes, including protein-protein interaction studies, cytokine detection, protein-DNA interaction studies, kinase target identification, antigen and antibody detection, and biomarker discovery. This method allows researchers to minimize the size of assays on a single platform. Protein microarrays enable comprehensive analysis of protein samples with minimal reagent consumption and labor-intensive procedures.
The Technology Behind Protein Microarrays
The process of creating a protein microarray involves several crucial steps, which include protein expression and purification, spotting the protein onto the array surface, and detecting protein interactions. Proteins of interest are usually produced using recombinant DNA technology or extracted from biological samples. They are then purified and printed onto the microarray substrate with robotic spotters.
Protein microarray slides
When printing proteins, researchers should consider the array’s surface type. Two common surfaces are epoxysilane-coated and nitrocellulose-coated glass slides, each with its own advantages and disadvantages.

Epoxysilane-coated slide

Nitrocellulose-coated slide
Epoxysilane-coated slides have minimal background fluorescence, which eliminates the need for a fluorescence subtraction process. However, this comes at the cost of a reduction in the amount of protein that can be bound per spot and a subsequent decline in signal intensity.
Nitrocellulose-coated slides, on the other hand, have a high capacity for protein binding, but they also exhibit significant background fluorescence that can vary across different samples.
Robotic microarray spotters
Manual microarray construction has inherent limitations, such as inaccuracy in spotting, a time-consuming process, reproducibility challenges, low throughput, and standardization difficulties. Robotic spotters address these issues by offering precise and consistent sample deposition, enhancing experiment reproducibility, increasing throughput, and aiding in standardization. These improvements significantly boost the quality and efficiency of microarray experiments. Aurora Biomed’s VERSA microarray spotter is a high-tech and high-precision automated liquid handling system that can be used in protein microarray preparation on various surfaces, including epoxysilane-coated glass slides, nitrocellulose-coated glass slides, membranes, micro-well plates, papers and biosensor substrates. In 2023, Krammer et al. conducted a study on a vaccine based on chimeric hemagglutinin (cHA). The researchers used Aurora Biomed’s microarray spotter to array avian influenza protein onto an epoxysilane-coated glass slide at the Icahn School of Medicine at Mount Sinai, New York. The purpose of this was to measure the breadth of antibody responses.
Protein microarray detection
There are two strategies used for protein microarray detection. The first is known as label-based detection. In this technique, researchers label query molecules with different tags, such as fluorescent dyes, epitope tags, and radioisotopes. This method is commonly used in protein microarrays because it is simple and reagents are widely available. However, these labelling strategies can alter the surface characteristics and natural activities of the query molecule. Additionally, the labelling process is time-consuming and laborious which restricts the types and number of query molecules that can be studied. However, researchers can use VERSA microarray spotter to simplify the microarray preparation process.
The second technique is known as label-free detection. In label-free techniques, researchers measure inherent properties of the query, such as mass or dielectric property. Label-free techniques avoid interference caused by tagging molecules and enable real-time determination of reaction kinetics in biomolecular interactions. However, label-free detection techniques also face challenges related to sensitivity and specificity.
Protein microarray types
Protein microarrays are available in different types, each serving its distinct purpose in proteomic analysis. The following are the most common types of protein microarrays:
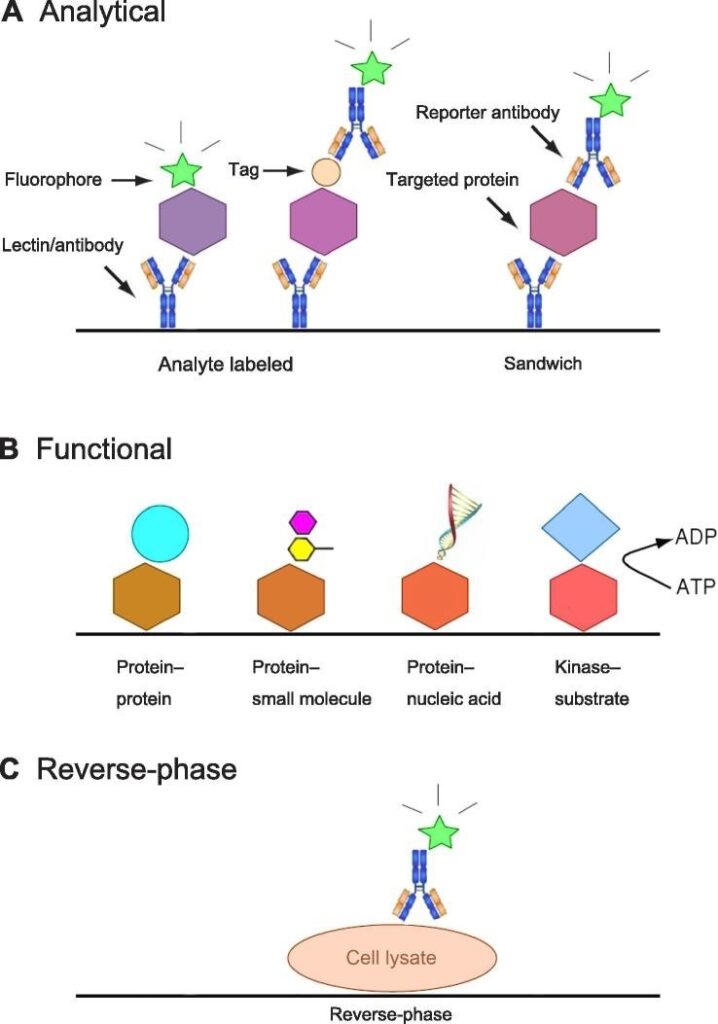
- Analytical Protein Microarrays:
These microarrays are the most commonly used and utilize antibodies as capture reagents. They can employ different assay formats, such as “analyte-labeled” or “sandwich” formats, to detect proteins. Though they offer high sensitivity and specificity, they pose certain challenges, such as nonspecific binding and the need for specific antibodies.
- Functional Protein Microarrays:
These protein microarrays are constructed using individually purified proteins. They enable studying biochemical properties, such as protein-protein interactions, enzyme-substrate relationships, and post-translational modifications. They have been instrumental in basic research and clinical applications, offering insights into proteomes of various organisms and post-translational modifications on a large scale.
- Reverse-Phase Protein Microarrays:
These microarrays enable the direct analysis of samples obtained at different states, such as tissue or cell lysates. They offer high sensitivity and precision, but their application is highly dependent on the availability and specificity of commercially produced antibodies.
Applications
Sim et al. (2012) developed a novel microarray-based method for detecting prostate-specific antigens using the VERSA microarray spotter. Ellebedy et al. (2020) utilized the VERSA 100 microarray spotter to create a microarray made of Influenza virus proteins. This microarray was used to detect specific antibody interactions, aiding in the study of immune responses to influenza virus proteins, which is important for research on influenza infection and vaccination.
Conclusion
In conclusion, protein microarrays offer a multifaceted approach to studying protein interactions and functions, serving as indispensable tools in proteomics research, clinical diagnostics, and drug discovery. With ongoing technological advancements, including innovations like the VERSA Microarray Spotter, these arrays continue to evolve, promising even greater insights into complex biological systems. As our understanding of protein biology deepens, protein microarrays stand poised to drive forward biomedical science, ultimately enhancing our ability to address critical health challenges.