
Automated Differential Digestion Analysis Utilizing DNase I
Advancing Justice for Survivors of Sexual Violence
Analyzing samples of sexual assault evidence kits (SAEK) requires using a precise technique called differential digestion to distinguish between sperm cells and non-sperm cells. As forensic laboratories faced the increase of public and legislative demands for timely examination of SAEK, automated liquid handling workstations were implemented to meet the demand. An automated differential digestion protocol was developed using a DNase I digestion step. The application of a degradative agent to selectively remove non-sperm DNA from mixed samples, allows automation of the differential digestion process, saving time and labor.
- No carryover of EC fractions
- Minimum Human Intervention
- ~5 hours for 96 samples
Independently Validated
A Decade of Automated Differential Digestions!
At ISHI34, Krina Carman and Monica Dyer from Oakland Police Department discussed their experience in processing 25000 Sexual Assault Evidence Kit (SAEK) samples with the help of VERSA 1100 in past 10 years.
Automation of a Differential Digestion Protocol for Processing SAEK using the VERSA Automated Liquid Handling Workstation
This method is currently employed by crime labs—Oakland PD has published a vigorous validation on the VERSA 1100. Follow the links below for more information and to view the latest publication from criminalist Helena Wong
Streamlining SAEK process at Contra Costa Sheriff’s Office with the VERSA 1100 DD Workstation
The Contra Costa County Office of the Sheriff faced a backlog of case works due to the number of cases coming in daily, especially with the complex protocol of processing SAEKs from victims. To address this issue, they turned to automation solutions and chose the VERSA 1100 Differential Digestion Workstation.
Overall Workflow
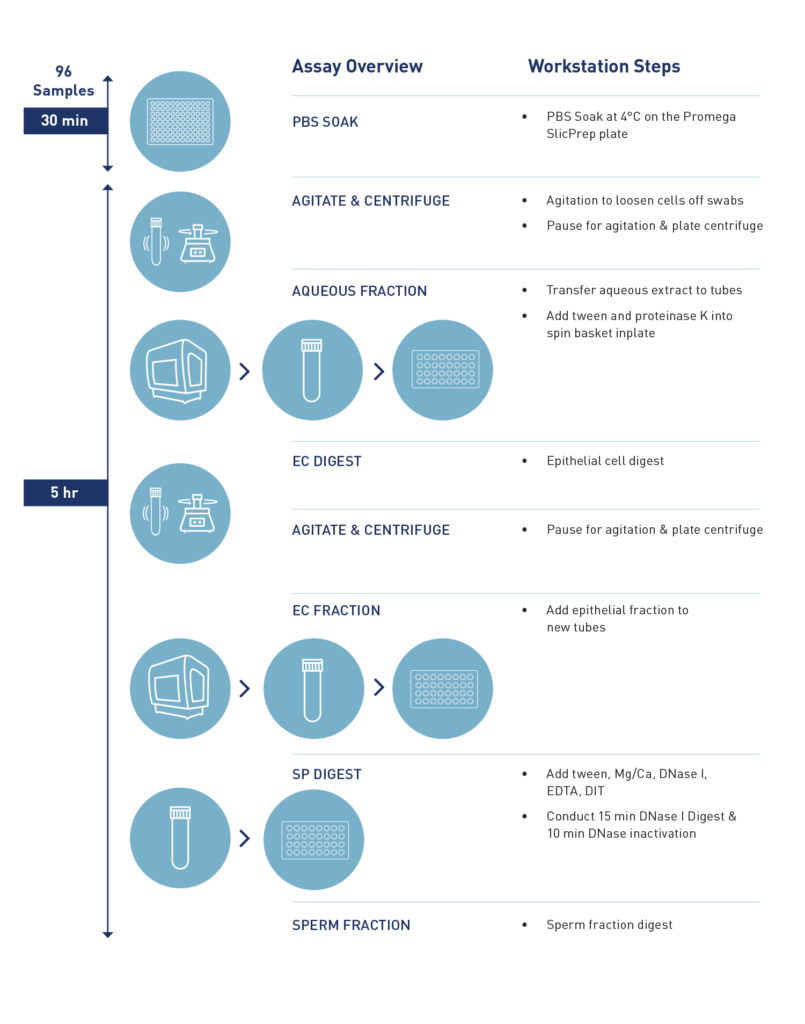
How long does it take to process 96 samples?
Processing a full plate of 96 samples typically takes about 5 hours, including incubation time on the workstation.
What is a brief overview of the automated protocol on VERSA?
The automated protocol starts by adding buffers to the spin basket, including samples, followed by a 30-minute rehydration period. Subsequently, a 30-minute shaking process loosens the swabs. The workflow pauses for the plate to be removed and centrifuged to pellet cells. Returning to the VERSA, it separates the supernatant into 2ml tubes. The process continues with the addition of digestion buffer and Pro-K, digesting epithelial cells for 30 minutes at 65 degrees on the on-deck heater. Following the removal of the epithelial cell fraction, the digestion targets sperm cells.
Does the VERSA1100 require specialized consumables or reagents for automating the differential digestion process?
Yes, the VERSA1100 requires specific automation consumables and reagents designed for differential digestion protocols, ensuring optimal performance.
Has the VERSA1100 differential digestion been validated for forensic samples?
Yes, the VERSA1100 differential digestion has been validated by a few forensic laboratories.
Can the purification step be carried out on the VERSA workstation?
Purification relies on magnetic bead separation and can be easily performed using the on-deck magnetic stand module.
How effective is VERSA for degraded samples?
A study evaluated various degraded sample types—exposed to multiple freeze-thaw cycles, high temperature, humidity, and aged semen stains. Results showed consistent DNA typing compared to traditional manual methods.
How is contamination addressed?
Contamination concerns were minimal due to VERSA’s built-in HEPA/UV system.
Can the VERSA automate slide preparation?
Yes, VERSA can automate both pre-digest and post-digest slide preparations.
Do the VERSA models have on-deck heaters?
Yes, VERSA is equipped with a Heater/Shaker on the deck, offering a temperature range of 4-90°C.
Can the manual agitation step be automated on the VERSA instrument?
Automating the manual agitation step poses significant challenges. Ensuring complete cell homogenization is crucial, but it’s difficult to achieve this without risking cross-contamination using the robotic arm. Concerns also arise regarding sample adherence to the pipette head, increasing the contamination risk. As a result, it was deemed more practical to retain this step as a manual process.





