On March 11th, 2020, COVID-19 developed into a blown pandemic threatening human health and the world’s economies. The disease, caused by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2), and for the first time was observed in Wuhan in December 2019. The COVID-19 virus was declared a pandemic by the World Health Organization (WHO). Since then researchers focused on learning more about this virus and how to encounter it.
It has been found that, a SARS-CoV-2 virus is a polyprotein that has four structural proteins (envelope, E; membrane, M; nucleocapsid phosphoprotein, N; spike, S), and five accessary proteins, Orf3a, Orf6, Orf7a, Orf8, and Orf10 (Figure 1). It also has been found that identifying B-cell and T-cell epitopes for SARS-CoV-2 proteins is an essential factor in developing practical diagnostic tests and vaccines.
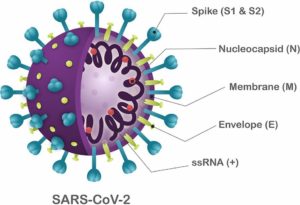
Figure 1 Structure of SARS-CoV-2.
Microarray, including cell microarray, protein microarray, and DNA/RNA microarray, all are considered as powerful tools in the ongoing research related to the COVID-19. These microarrays can analyze hundreds or even thousands of samples in a single run, making them much faster than traditional virus detection methods. These techniques allow researchers to rapidly screen for the presence of viral proteins in samples, providing valuable information about the virus and its behavior.
Regarding the protein microarray, one of its key benefits is its high throughput nature. This is particularly useful in the early stages of an outbreak when rapid identification of infected individuals is critical to contain the spread of the virus.
Another advantage of protein microarrays is their ability to detect multiple proteins simultaneously. Traditional virus detection methods often require multiple tests to identify the presence of different viral proteins, but protein microarrays can detect multiple proteins in a single assay. This can help researchers understand how the virus behaves and evolves over time.
In addition to their use in virus detection, protein microarrays are also used to study the immune response to SARS-CoV-2. By analyzing the proteins in blood samples from infected individuals, researchers can learn more about how the body responds to the virus and how it fights against the infection. This information is critical to developing effective treatments and vaccines for the virus. And overall, protein microarrays are a priceless tool in SARS-CoV-2 research..
One such an example of using peptide microarrays is from Dr. McGeer research group from the University of British Columbia (UBC). They have prepared peptide chips to detect antibodies against the SARS coronavirus in serum (Figure 2). Another example is the research at Beijing Proteome Research Center, who developed a SARS-CoV-2 proteome peptide microarray to perform the proteome-wide analysis of serum antibody’s epitopes at amino acid resolution.
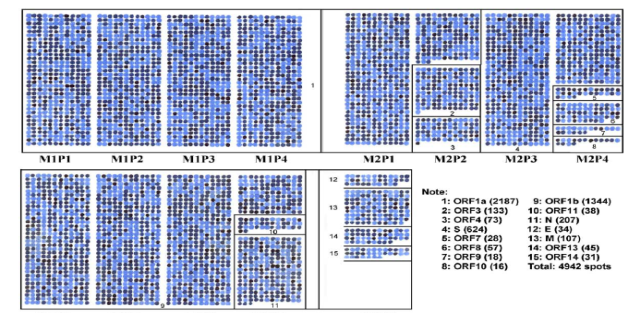

Figure 2 Outline of the overlapping peptide set covering membranes M1, M2, M3, and M4.
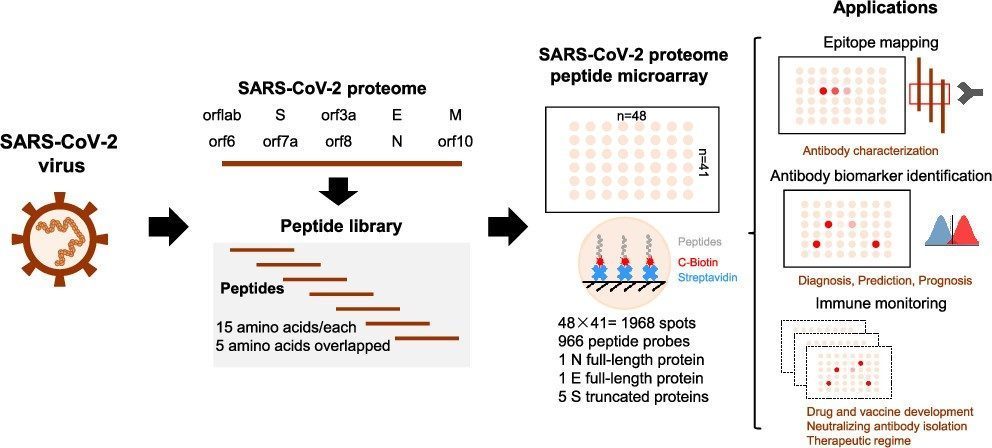
Figure 3 The schematic illustration of SARS-CoV-2 proteome microarray fabrication and biomedical applications.
In this regard Aurora Biomed has developed a peptide microarray instrument known as VERSA Spotter. Aurora Biomed’s VERSA automated peptide microarray spotter has been successfully used to develop the Omicron S protein microarray (Figure 4). The protein microarray was prepared by the solid-phase synthesis method, consisting of 421 peptides. Each peptide is 15 amino acids (AA) in length, with an overlap of 12 amino acids between two adjacent peptides.

Figure 4 COVID-19 Variant Omicron S protein microarray.
The Versa Peptide Microarray Spotter workstation can streamline workflows in epitope mapping, biomarker identification, enzyme-substrate profiling, and drug development. The NanoPipettor is capable of repeatedly pipetting sub-microliter amino acid volumes for synthesizing high-density microarrays in simple or complex user-defined patterns. The customizable deck layout support printing on diverse substrates, including filter papers



