In the realm of cancer diagnosis, precision and efficiency are two important factors. The advancement of technology has opened doors to groundbreaking methodologies that have revolutionized the field. One such innovation making an insightful impact is the automated microarray spotter. This cutting-edge instrument revolutionizes how we detect and analyze genetic information, offering unparalleled comprehensions for accurate cancer diagnosis. In this blog post, we will investigate the world of microarrays and explore how automation solves the potential of microarray, driving the future of precision medicine.
Microarrays have changed the landscape of cancer diagnostics by unraveling the intricate genetic makeup of cancer cells. By simultaneously analyzing the expression of thousands of genes, microarrays provide a comprehensive understanding of the molecular underpinnings of various cancers. This information allows precise characterization prognosis assessment, and personalized treatment selection.
DNA microarray is one of the main tools based on molecular biology to analyze gene expression. DNA microarray, also known as gene chip or DNA chip, is a glass slide or silicon substrate containing thousands to millions of tiny spots called probes. Probes are short DNA sequences complementary to specific genes or regions of interest in the genome.
Cancer tissue or cells are collected from patients and processed to extract RNA, representing the genetic material that carries the information for gene expression (Figure 1). The RNA is reverse-transcribed into complementary DNA (cDNA) and then labeled with fluorescent dyes. The labeled cDNA from the patient’s sample is mixed with the DNA microarray. The cDNA binds to the complementary probes on the microarray through hybridization. The hybridization occurs when the sequences of the cDNA and probes are complementary, resulting in the formation of cDNA-probe pairs. The microarray slide is scanned using a laser or other suitable imaging system that detects the fluorescent signals emitted by the labeled cDNA. Each spot on the microarray corresponds to a specific gene or genomic region, and the intensity of the fluorescence signal represents the gene expression level. The fluorescence intensity data obtained from the microarray image is processed and analyzed using bioinformatics tools and software. The analysis involves comparing the expression levels of different genes between cancerous and normal tissues or between different cancer subtypes. Statistical methods are applied to identify significantly upregulated or downregulated genes in cancer cells. Based on the gene expression patterns identified through microarray analysis, clinicians and researchers can gain insights into the molecular characteristics of cancer, classify tumors into different subtypes, predict prognosis, and guide treatment decisions. The identified gene signatures can serve as potential biomarkers for cancer diagnosis, prognosis assessment, and the development of targeted therapies. DNA microarray technology provides a comprehensive view of the molecular alterations associated with cancer by analyzing the gene expression profiles in cancer cells.
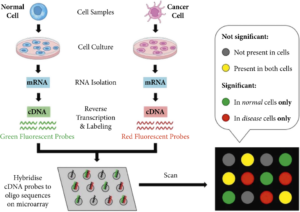
Figure 1. Workflow of DNA Microarray for Monitoring Gene Expression of Cancer Cells.
Cell microarray is another type of microarray that can be used for cancer diagnosis. It involves immobilizing cells on a solid substrate in a spatially organized manner, allowing for high-throughput analysis of cellular behavior, molecular profiles, and responses to therapeutic agents (Figure 2). Cell microarray allows researchers to analyze the expression levels of multiple biomarkers in cancer cells. By staining the microarrayed cells with specific antibodies or fluorescent probes, researchers can identify and quantify the presence of specific proteins or genetic markers associated with cancer. This information can aid in the diagnosis, classification, and prognosis of different types of cancer. Also, cell microarray facilitates assessing cancer cell response to various anti-cancer drugs or therapeutic agents. By treating the microarrayed cells with different compounds, researchers can evaluate the sensitivity or resistance of cancer cells to specific drugs. This information helps guide treatment decisions and the development of personalized therapeutic approaches.
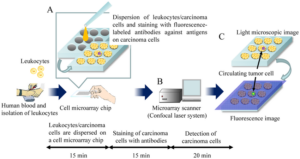
Figure 2. Schematics of Cell Microarray for Detection of Circulating Tumor Cells.
Yet, the manual construction of microarray has inherent limitations, such as inaccuracy in spotting, time-consuming process, reproducibility challenges, low throughput, and standardization difficulties. To overcome these limitations and unravel the full potential of microarray technology in cancer diagnosis, the integration of automation has become essential. Aurora VERSA Microarray Spotter solves these challenges by offering precise and consistent spotting, increased throughput, and improved reproducibility. A research team at National Taiwan University used our VERSA Microarray Spotter to develope a novel nanodroplet-based cell processing platform designed to evaluate drug synergy in anti-cancer treatments. The nanodroplet cell processing platform offers a promising approach to studying drug combinations, potentially leading to improved cancer treatments by identifying effective synergistic drug pairs. With its unparalleled precision, high-throughput efficiency, versatility, user-friendly software, and streamlined automation, the VERSA Microarray Spotter empowers researchers worldwide to reach new heights in cancer studies.



